Project members
Project
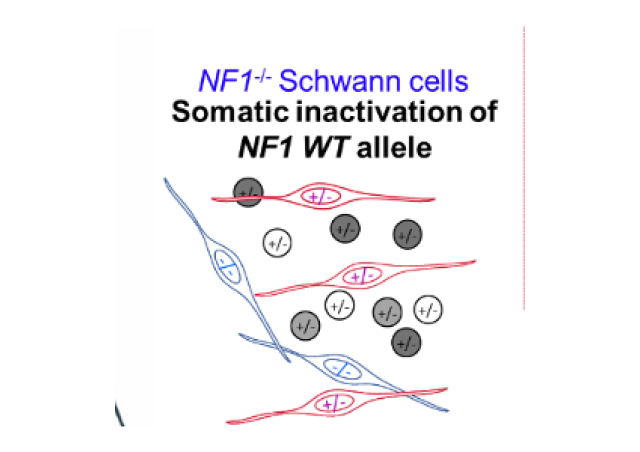
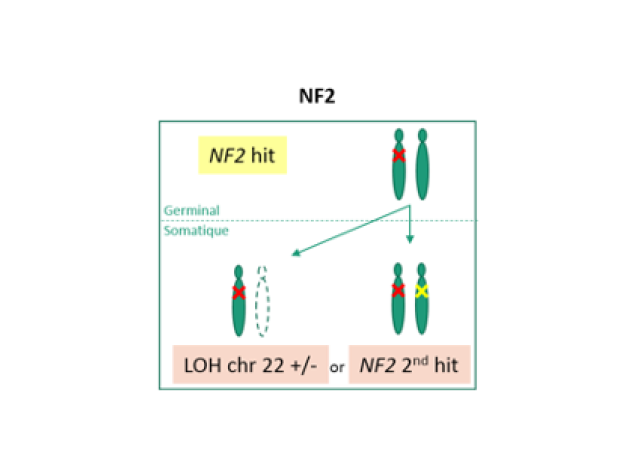
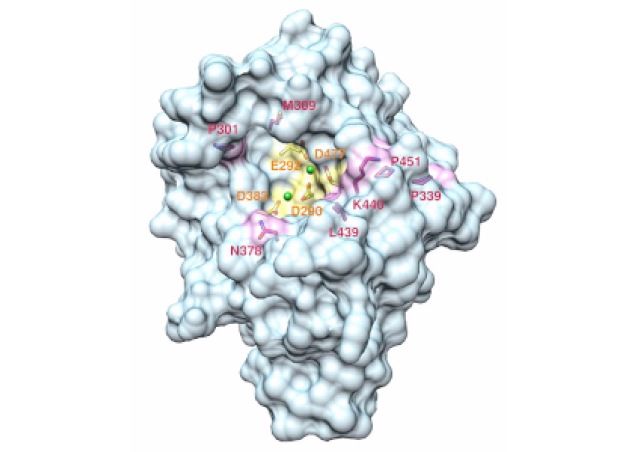
NF1 is a major tumor suppressor gene encoding neurofibromin, a RAS-GAP (GTPase activating protein) that acts as an inhibitor of the RAS-MAPK pathway by allowing RAS proteins to return to inactive confirmation. NF1 gene alterations and their therapeutic targeting have been little studied in sporadic cancers because NF1 is a long gene with many pseudogenes and no mutation hot spot.
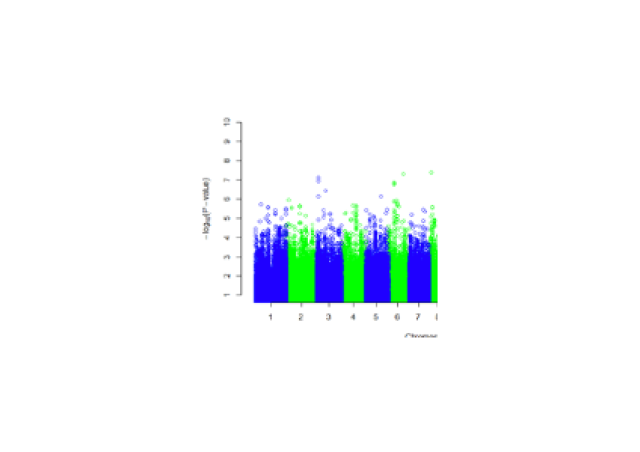
We have characterized genetic alterations in the NF1 gene in a cohort of non-small cell lung carcinomas (NSCLC) using a targeted NGS approach to confirm NF1 involvement and to characterize genotype/phenotype correlations. A NF1 genetic alteration was found in 24/137 (17%) samples (Tlemsani et al., 2019). Our objectives are to understand the functional consequences of NF1 inactivation and to identify new potential therapeutic targets in NF1-mutated bronchial carcinomas which represent a distinct subtype of cancer. We have developed cellular models of NF1-mutated bronchial carcinoma, using the CRISPR Cas9 genome editing technique. Our hypothesis is that NF1 mutations in the context of NSCLC carcinogenesis could cause cellular vulnerabilities that constitute therapeutic targets by synthetic lethality. We explore these synthetic lethalities with NF1 mutations using pharmacological and genetic approaches
In addition to bronchial tumors, our team is also interested in the characterization of NF1 mutations and more broadly of negative regulators of the RAS-MAPK pathway in pediatric acute lymphoblastic B leukemia (ALL-B). The activation of the main effectors of the pathway is well described but the alterations of the regulators are still poorly understood. Our project is to characterize (1) the transcriptome variations of a panel of negative regulators of the pathway, using a targeted transcriptomic study, and (2) the genetic alterations of these effectors in a large cohort of pediatric B-ALLs (including matched relapse samples), using a targeted NGS sequencing study.