Project members
Project
The overall goal of the project is to define the regulatory mechanisms determining the myofiber type of adult skeletal muscle and more specifically the mechanisms controlling the expression of a single myosin heavy chain (Myh) gene in the hundreds of nuclei present in each fiber. The major myofiber types are generally defined by the Myh genes that they expressed in the fiber. However, we lack comprehension of the regulatory mechanisms that determine which Myh gene will be expressed within a myofiber. The fast Myh locus in mammals is composed of Myh2, Myh1, Myh4, Myh8 and Myh13 genes arranged in 300Kb. Previous work from our group has identified by 4C experiments a 42kb super enhancer (SE) upstream of Myh2, that can interact with the promoters of Myh2 in soleus and with the promoter of Myh4 in quadriceps. This SE is composed of discrete DNA elements as revealed by single nucleus ATAC-seq (snATAC-seq) experiments performed with adult skeletal muscles suggesting that this SE could control fast Myh genes expression. Our research will characterize how the fast subtype myofiber gene expression program is established. In particular we will test the hypothesis that the SE at the Myh locus is responsible for organizing alternate DNA conformations to ensure that only a single Myh gene is expressed in the muscle fiber and the role of fast subtypes motoneurons in its activity.
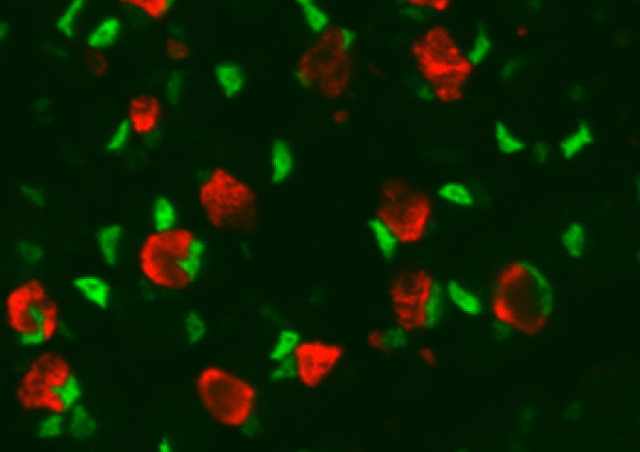
To carry out our program, we have implemented several complementary approaches including unique tools using BAC-Myh transgenic mice, CrispR/Cas9 KO models, snATAC-seq and snRNA-seq on adult muscles and RNAscope on isolated myofibers experiments in order to perform fiber-type specific genomic analyses on in vivo-derived muscle to understand the basic mechanisms defining myofiber fast subtype specification, notably apprehend why all myonuclei within a given myofiber express a unique Myh fast gene while the adjacent genes are silent. The role of the identified SE at the locus and the influence of motoneurons to specify its activity will be analyzed.

The fast Myosin heavy chain super enhancer (fMyh-SE) is composed of seven enhancer elements (1-7) recruiting TF and cofactors allowing the nuclear formation of a phase separation condensate in myonuclei and allowing robust fMyh expression. In the model of competition/exclusion, Myh2, Myh1, Myh8 and Myh13 promoters are excluded from the phase-separated droplet in Myh4+ myonuclei because they are not bound by sufficient amount of TF and cofactors. In those myonuclei robust bi-allelic expression of Myh4 is achieved, while transcription of the other genes in not detected. The model is not to scale.